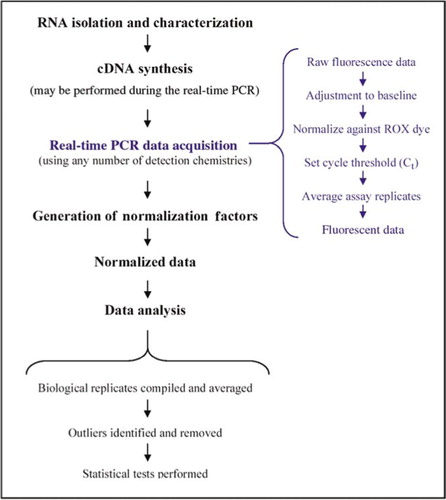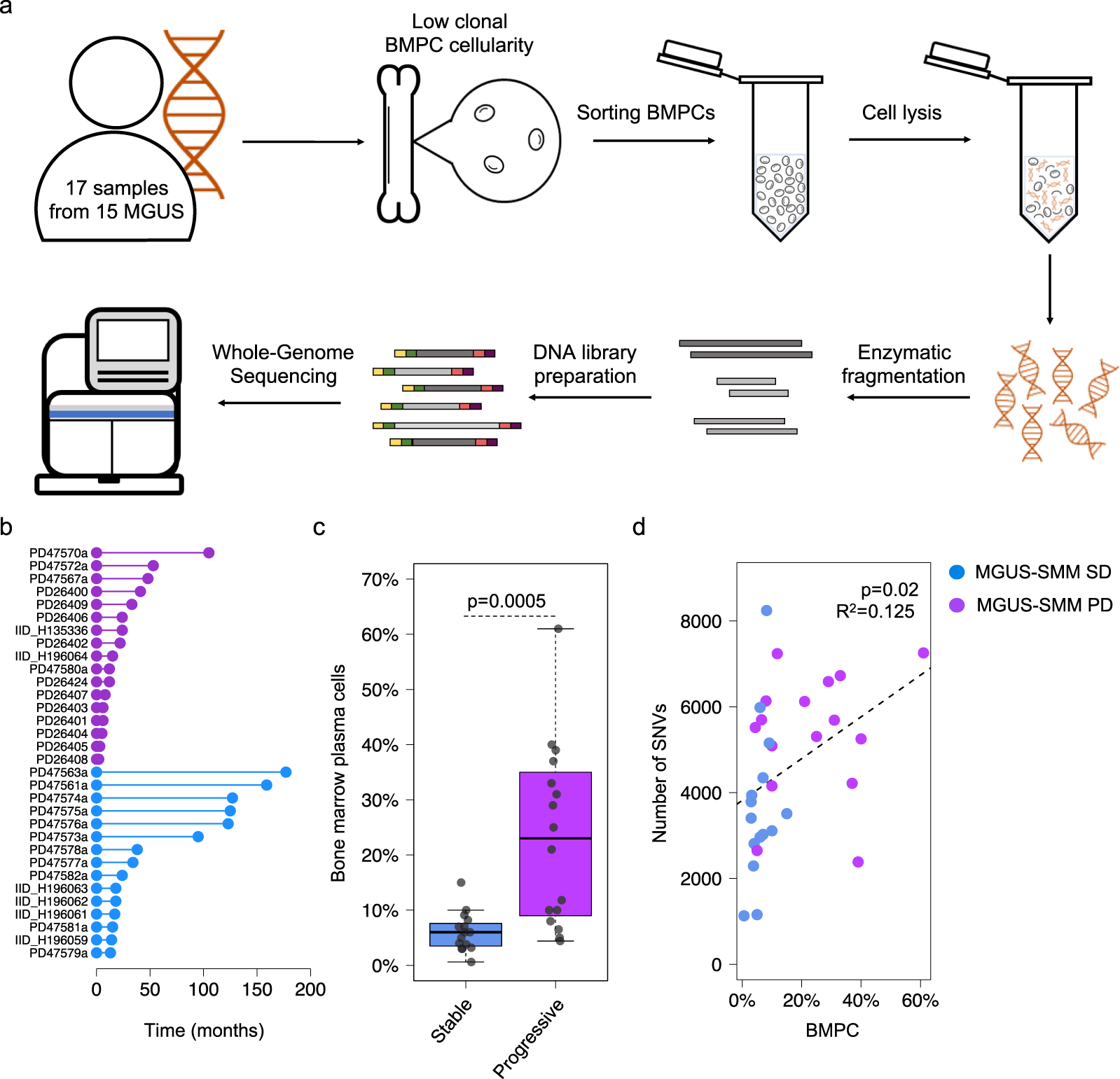
Whole-genome sequencing reveals progressive versus stable myeloma precursor conditions as two distinct entities | Nature Communications
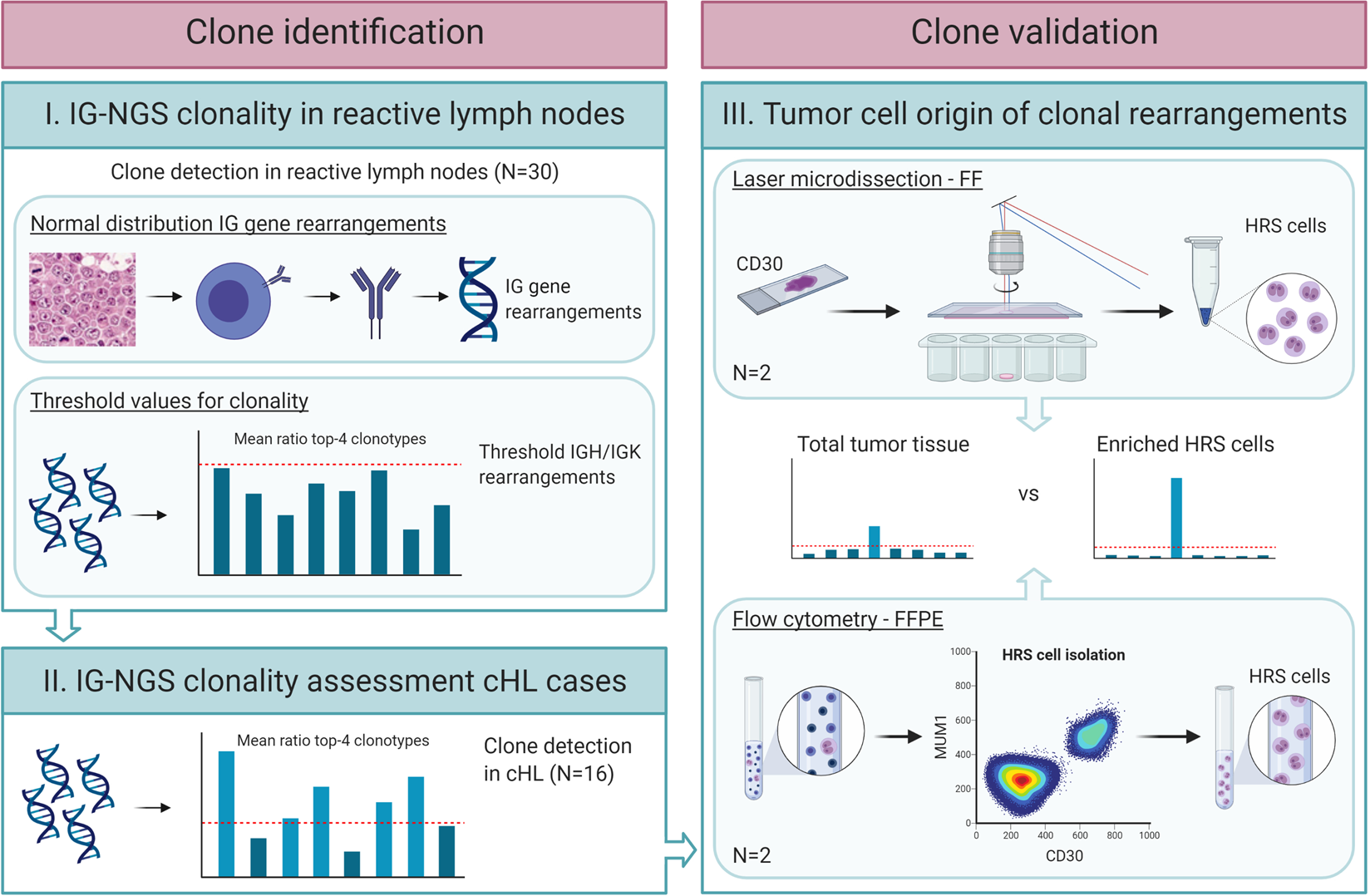
Clonality assessment and detection of clonal diversity in classic Hodgkin lymphoma by next-generation sequencing of immunoglobulin gene rearrangements | Modern Pathology
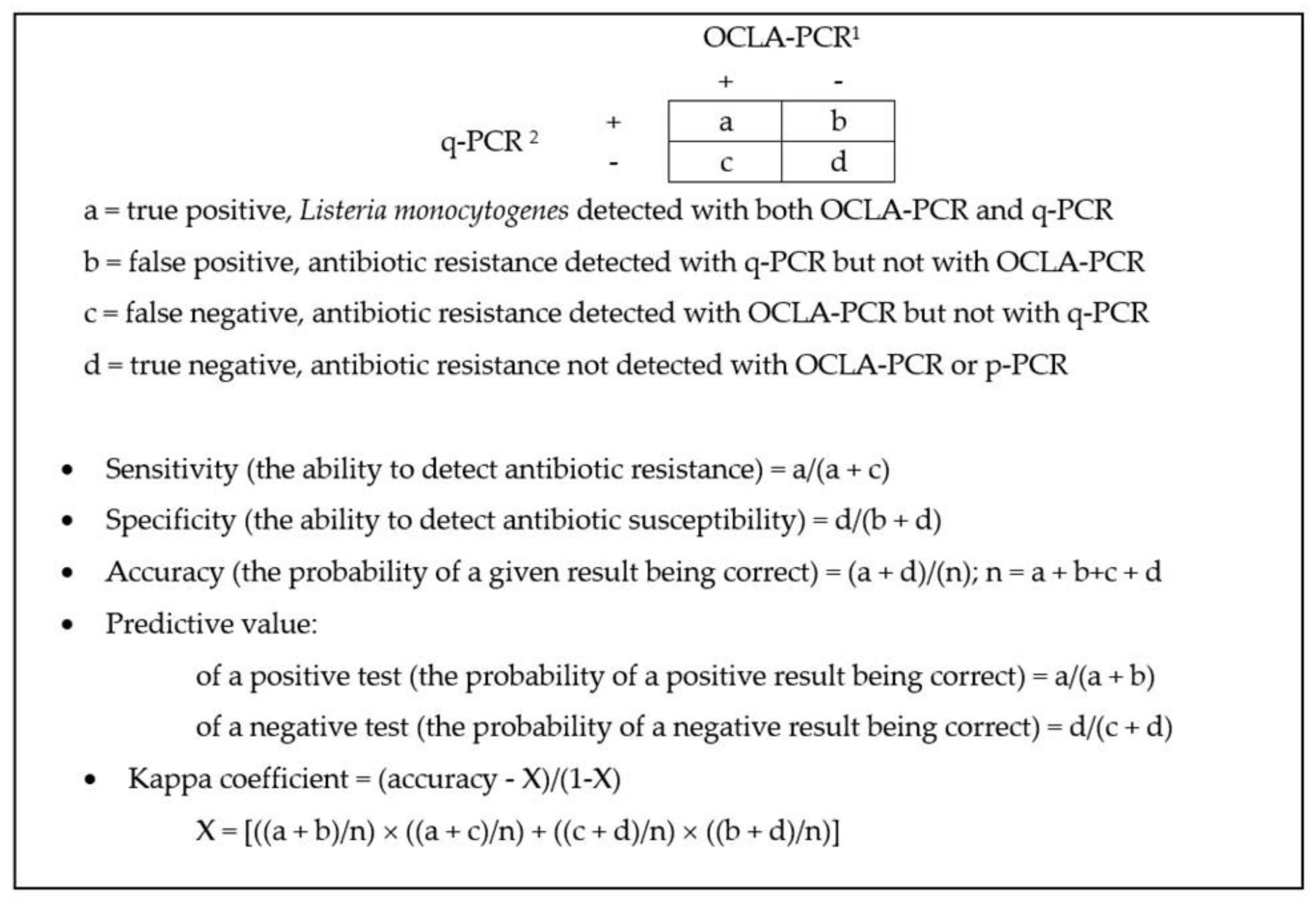

Antibiotics | Free Full-Text | Quantification of Total and Viable Cells and Determination of Serogroups and Antibiotic Resistance Patterns of Listeria monocytogenes in Chicken Meat from the North-Western Iberian Peninsula

Diagnostic performance of a colorimetric RT -LAMP for the identification of SARS-CoV-2: A multicenter prospective clinical evaluation in sub-Saharan Africa - eClinicalMedicine
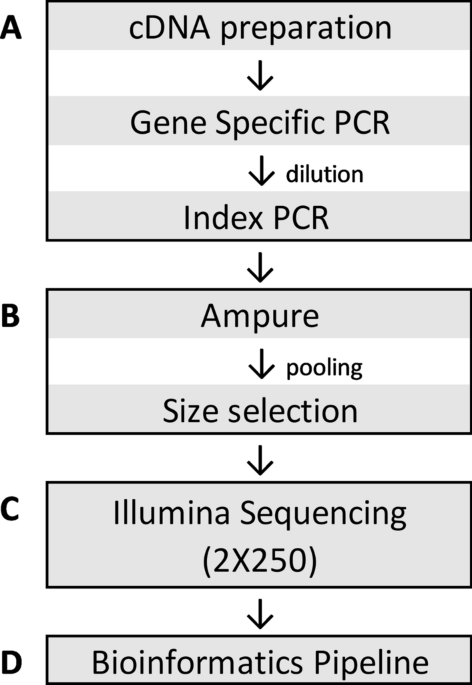
Development and validation of a high throughput SARS-CoV-2 whole genome sequencing workflow in a clinical laboratory | Scientific Reports
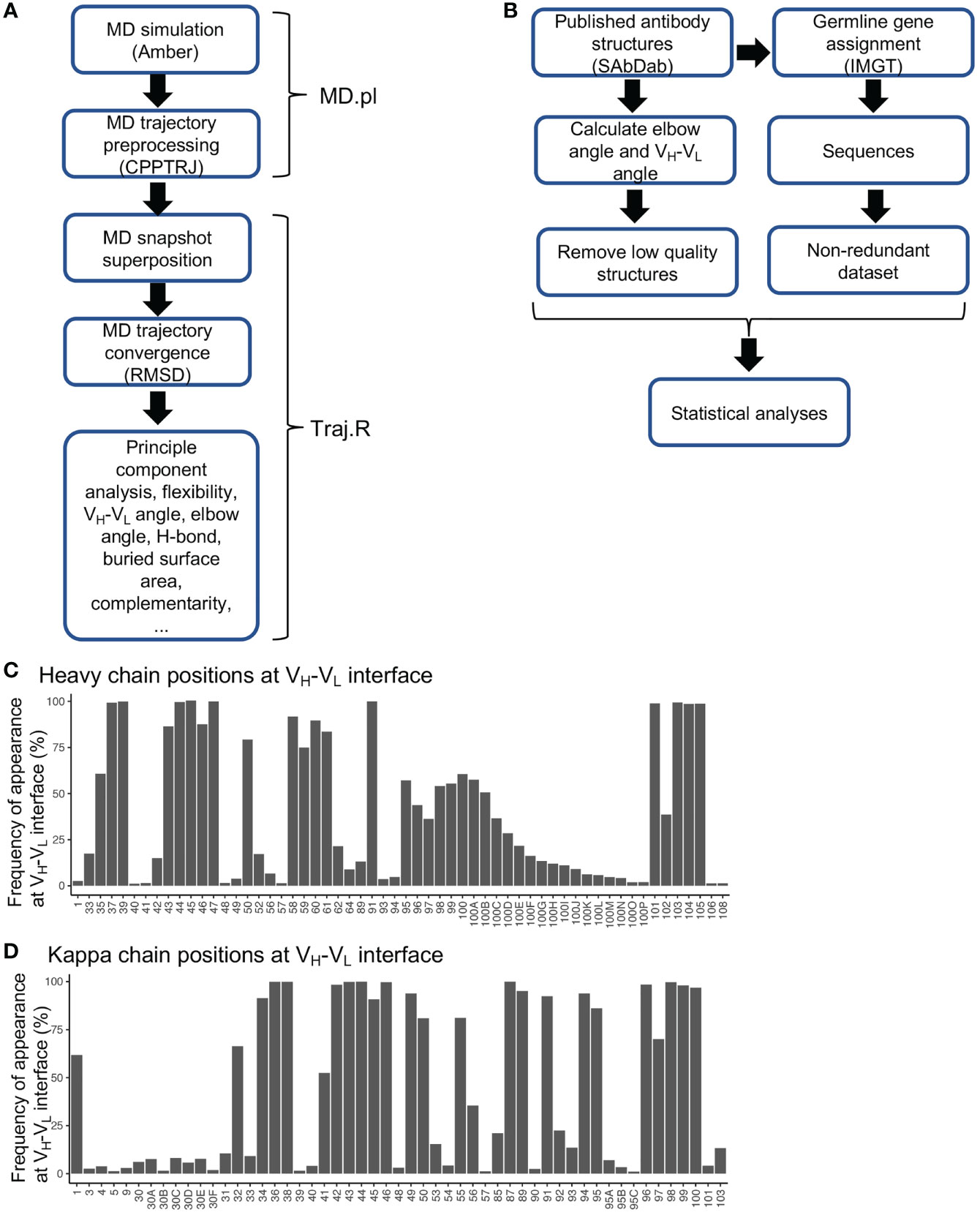
Frontiers | Structural Basis of Antibody Conformation and Stability Modulation by Framework Somatic Hypermutation

Unreacted Labeled PCR Primers Inhibit the Signal in a Nucleic Acid Lateral Flow Assay as Elucidated by a Transport Reaction Model | ACS Measurement Science Au

Novel One-Step Single-Tube Nested Quantitative Real-Time PCR Assay for Highly Sensitive Detection of SARS-CoV-2 | Analytical Chemistry

Paired heavy- and light-chain signatures contribute to potent SARS-CoV-2 neutralization in public antibody responses - ScienceDirect
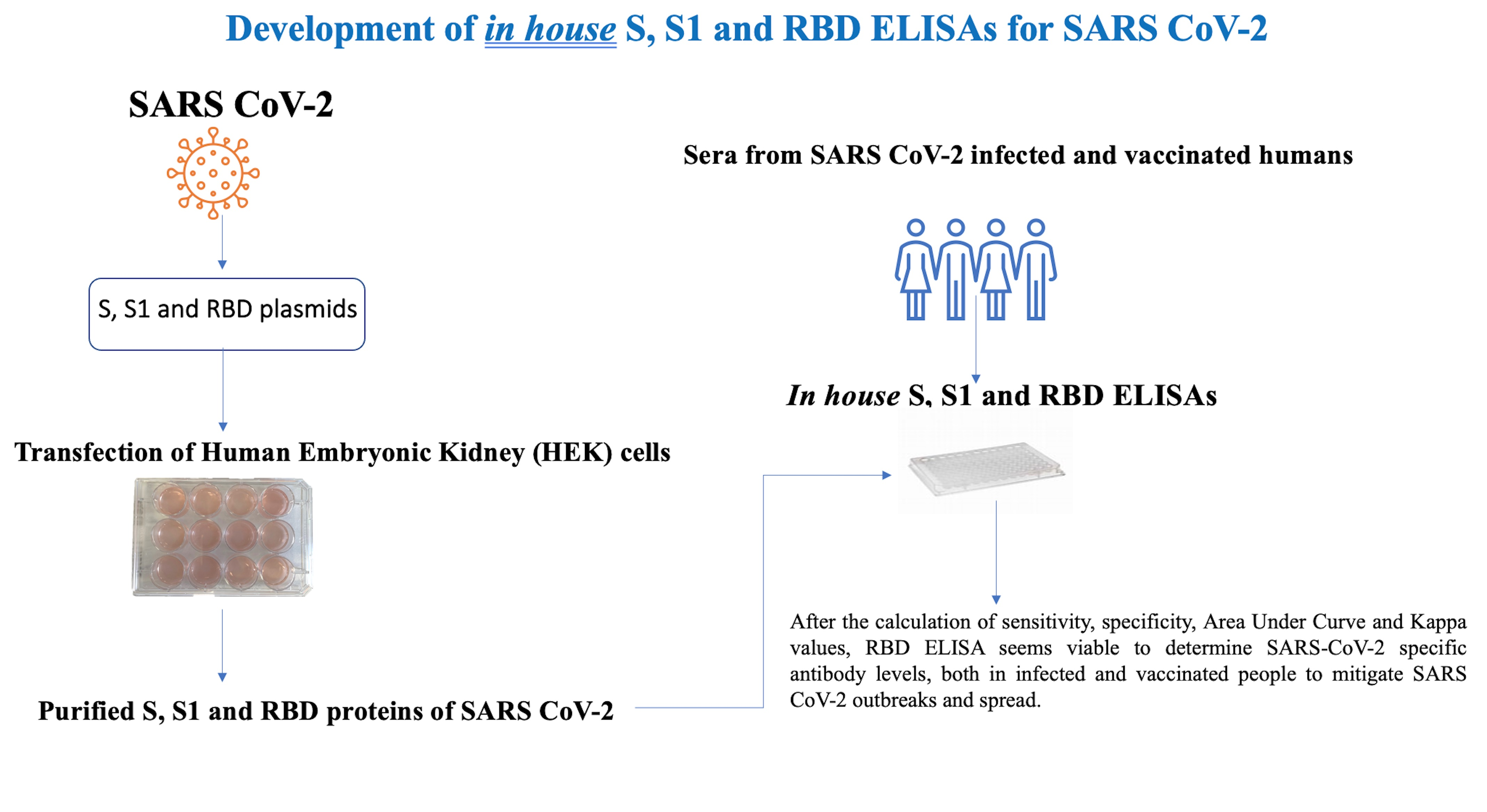
Diagnostics | Free Full-Text | Development of in House ELISAs to Detect Antibodies to SARS-CoV-2 in Infected and Vaccinated Humans by Using Recombinant S, S1 and RBD Proteins
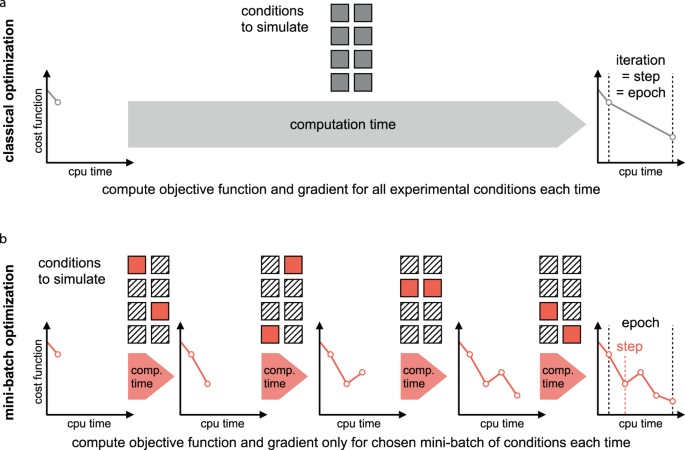
Mini-batch optimization enables training of ODE models on large-scale datasets | Nature Communications
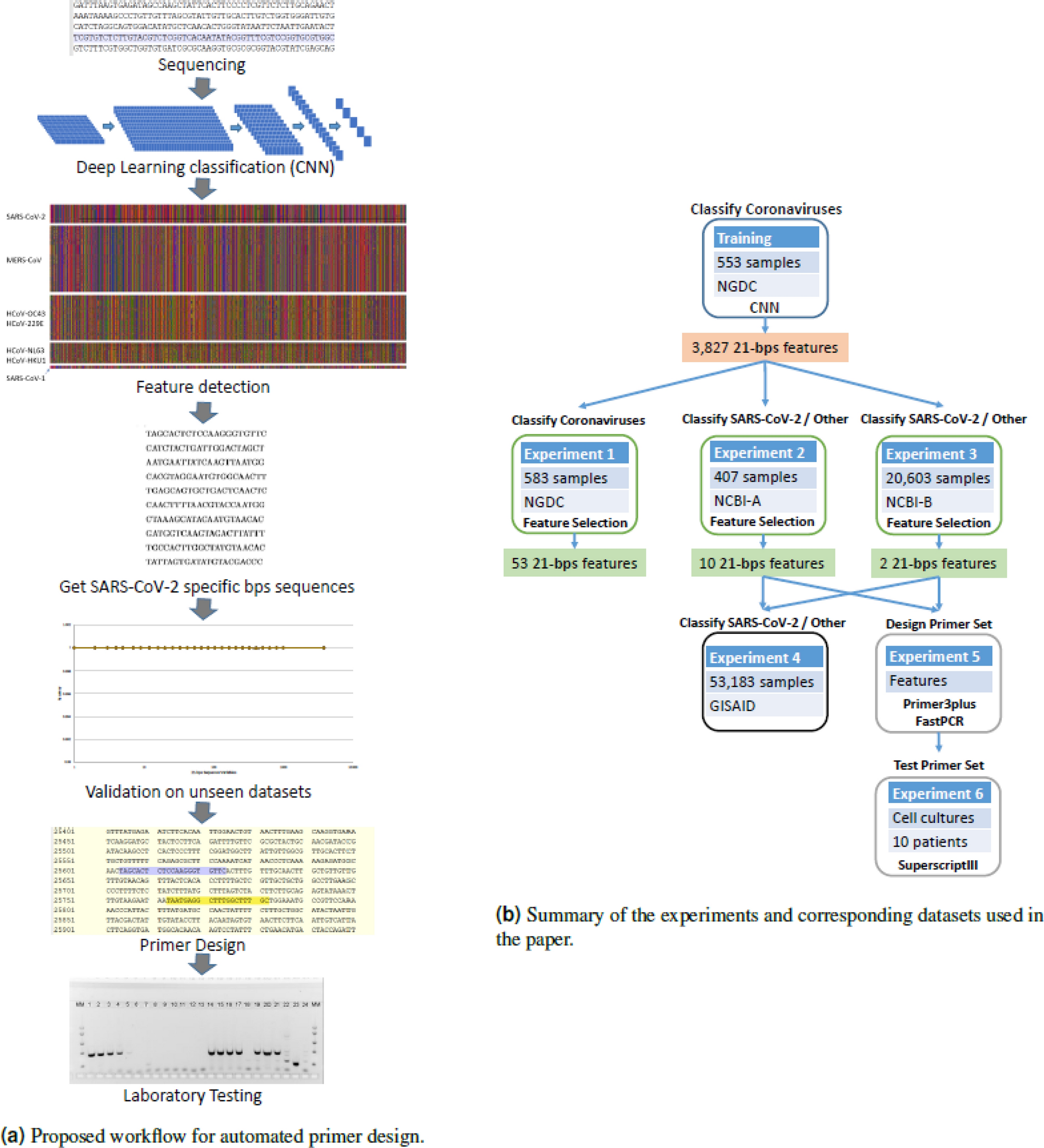
Classification and specific primer design for accurate detection of SARS-CoV-2 using deep learning | Scientific Reports

Validation of a rapid antigen test as a screening tool for SARS-CoV-2 infection in asymptomatic populations. Sensitivity, specificity and predictive values - eClinicalMedicine
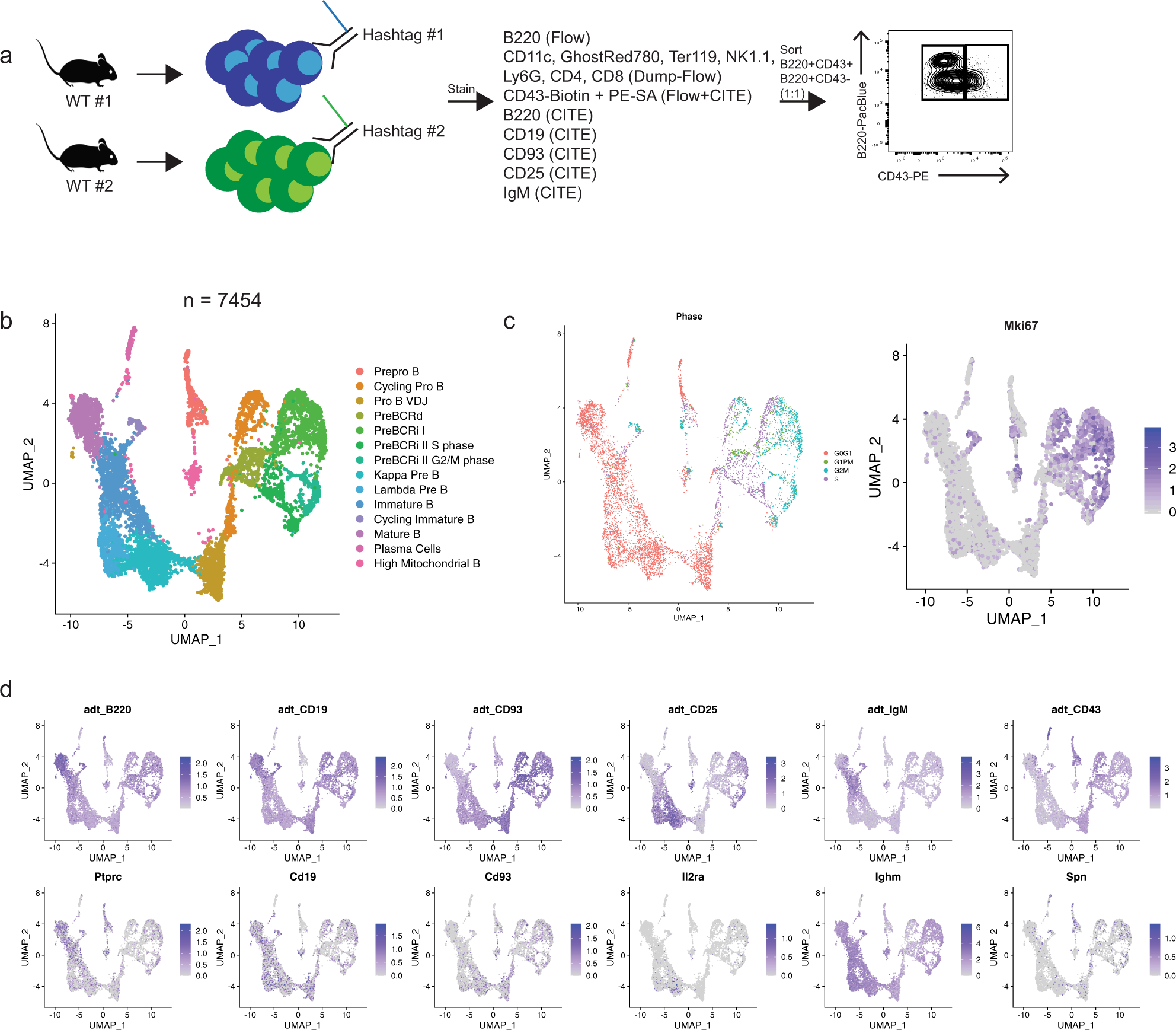
Single-cell analysis identifies dynamic gene expression networks that govern B cell development and transformation | Nature Communications

Comparison of Real‐Q 2019‐nCoV and DaAn Gene 2019‐nCoV polymerase chain reaction assays for the detection of SARS‐CoV‐2 - Marembo - 2022 - Journal of Clinical Laboratory Analysis - Wiley Online Library

When the Dust Has Settled: Calculation of Binding Affinities from First Principles for SARS-CoV-2 Variants with Quantitative Accuracy | Journal of Chemical Theory and Computation
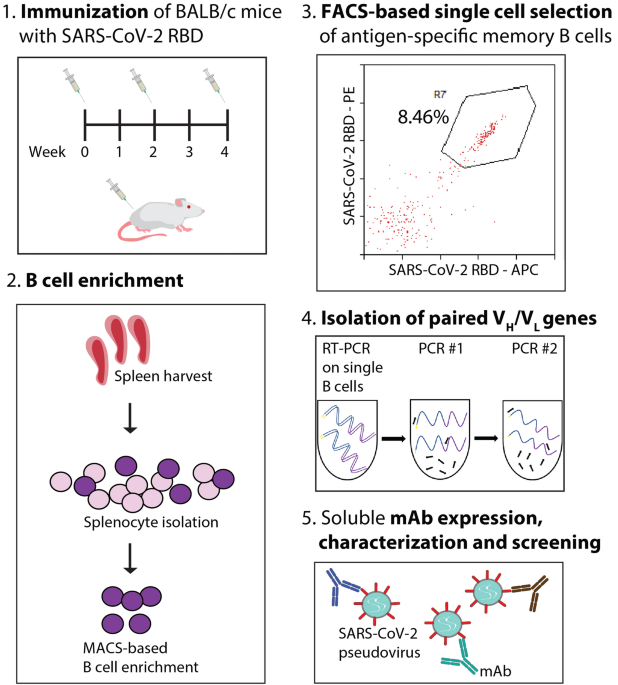
Discovery and characterization of high-affinity, potent SARS-CoV-2 neutralizing antibodies via single B cell screening | Scientific Reports
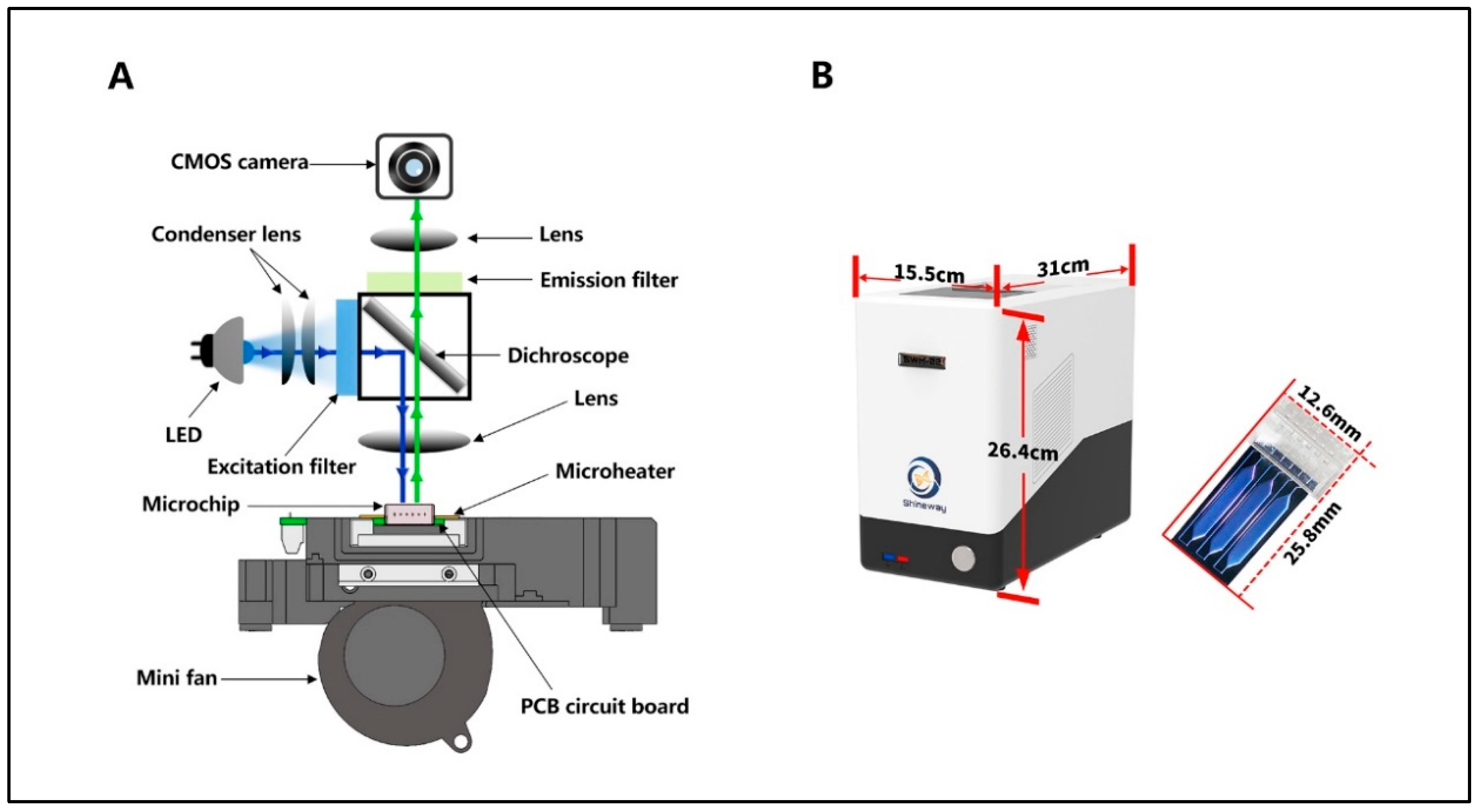
Bioengineering | Free Full-Text | Ultrafast PCR Detection of COVID-19 by Using a Microfluidic Chip-Based System
Diagnostic accuracy and acceptability of molecular diagnosis of COVID-19 on saliva samples relative to nasopharyngeal swabs in tropical hospital and extra-hospital contexts: The COVISAL study | PLOS ONE